Editorial
ZEB1 is an Oncogenic Factor that Represses Many Tumor Suppressors in Uveal Melanoma
Yao Chen1,2, Ling Gao1* and Yongqing Liu2*
1Second Affiliated Hospital, Central South University Xiangya School of Medicine, Changsha, China
2Department of Ophthalmology and Visual Sciences, University of Louisville, Louisville, USA
*Corresponding author: Yongqing Liu, Department of Ophthalmology and Visual Sciences, University of Louisville, 301 E. Muhammad Ali Blvd., Louisville, KY 40202, USA
Published: 03 Apr, 2017
Cite this article as: Chen Y, Gao L, Liu Y. ZEB1 is an
Oncogenic Factor that Represses Many
Tumor Suppressorsin Uveal Melanoma.
Clin Oncol. 2017; 2: 1256.
Editorial
Uveal Melanoma (UM) is the most common intraocular tumor in adults, with an incidence
rate of 4.3 per million in USA [1,2]. Half of the patients developdeadly metastasis preferentially to
the liver and no effective treatment is available for the metastatic lesion [3,4]. UMsare transformed
from the melanocytes ofneural crest originin the uveal tractwith a mobile property [5] and
pathologically classified into spindle (mesenchymal-like), epithelioid (epithelial-like) and mixed
cell types [6] with a tendency of the epithelioid tumorsto develop a metastatic disease [7]. Abnormal
expression of some genes are found in UM transformation and progression including ZEB1. ZEB1
is a member of theE-box binding Transcription Factor (TF) group that includes ZEB, SNAI, and
TWIST familiesfunctioning in Epithelial to Mesenchymal Transition (EMT) in carcinomagenesisin
whichnormal epithelial cells are transformed andgain cell flexibility in escaping their primary
sites [8,9] and acquisition of cancer stem cell features likeresistance of conventional radio- and
chemotherapies, disease recurrence and poor prognosis [10]. In contrast, UM is a non-epithelium
tumorso thatdoes not necessarily proceed through EMT for transformation and progression as the
melanocyte-derived tumor has already acquired mesenchymal propertiesregardless of their cell
morphology. In fact, the spindle UM is pathologically consideredless aggressive than the epithelioid
UM though UM aggressiveness is still positively related to high expression of the EMT-TFs [6,11,12],
suggesting that the mechanism underlying EMT-TF regulation of tumorigenesis is not through
morphological EMT per se, but due to their molecular involvement in cell mitosis, mobility, and
adaptability behind the morphological switch.Here, we desire to briefly assess how ZEB1 is involved
in regulation of UM progression based on our recent investigation [11].
Our laboratories and others’ have revealed that ZEB1 is almost undetectable in Normal Uveal
Melanocytes (NUM), but states high in primary UM and even higher in Metastasized UM (MetUM),
and that its expression is significantly higher in epithelioid cells than in spindle cells [11], suggesting
that ZEB1 involves in UM tumorigenesis and malignant evolution. We therefore reason that initial
transformationofelongated fibroblast-likeNUM to the spindle UM and then development to the
epithelioid UM phenotypeis delimited bya gradually increased ZEB1 expression.This assumption
was experimentally verified by ourobservations both in vivowhere the spindle OCM1 UM cell linegrafted
tumors displayed both spindle and epithelioid cell types while the epithelioid C918 UM
cell line-grafted tumors only gave rise to epithelioid cell type, and in vitro wherea single spindle
OCM1 cell-formed organoidcould generate cells of various shapes including spindle and epithelioid
phenotypes while a single epithelioid C918 cell-formed organoidcould only produce epithelioid
cells [11]. Furthermore, overexpression of ZEB1 in ZEB1low OCM1 and knockdown of ZEB1 in
ZEB1high C918 did not affect their cell morphology [11], suggesting that ZEB1 has little effect on UM
cell morphology, but is a major oncogenic factor for UM progression.
To assessbiological functions of ZEB1 in regulation of UM progression in detail, we adopted a
loss-of-function approach using shRNAto knockdown ZEB1 expression in both OCM1 and C918
UM cell lines for grafting in the vitreous of the nude mice and for molecular analyses [11]. As results,
we find that knockdown of ZEB1 significantly reduces UM cell proliferation both in culture and in
the grafted tumors. The cause of ZEB1-induced cell proliferation is linked tothe binding of ZEB1
to a group of Cyclin-Dependent Kinase Inhibitors (CDKIs) such as P21 that prevent activation of
RB1, amajor tumor suppressor and cell cycle regulator [13]. More importantly, ZEB1 is observed
to bind to and repress another cell cycle regulator and also a well-known UM suppressor BRCA1
associated protein 1 (BAP1) [11,14]. Secondly, we find that ZEB1 highly expresses in non- or less
pigmented UM cells as it binds to the melanocyte promoter of the pigment synthesis regulator gene MITF and repress its expression. We link the ZEB1-repressed
MITF expression to the dedifferentiated state of UM, indicative of
a more advanced stage. And again, ZEB1 is observed to bind and
repress another cell differentiation facilitator and also a well-known
UM suppressor ID2 (inhibitor of DNA binding protein 2) [11,15]. It
is of note that the famous carcinoma suppressor E-cadherin (CDH1)
is hardly detectable in most primary UMs [11,15] and cell lines, and
knockdown of ZEB1 has little effect on its expression [11], suggesting
that ZEB1 mechanistic regulation of tumorigenesis in UM differs from
carcinomas. Thirdly, knockdown of ZEB1 degrades the invasiveness
of grafted UMs in the vitreous of the model animals likely by
reducing tumor cell capacity to break through the healthy tissues as
ZEB1 can bind to and thereby transactivate thoseextracellular matrix
degradation enzyme genes likematrix metalloproteinase 11 (MMP11)
and urokinase-plasminogen activator (PLAU) [11]. Fourthly,
knockdown of ZEB1 significantly decreases mobility of UM cells
in culture and reduces UM metastasis in the grafted model animals
[11]. ZEB1-increased UM cell mobility is accomplished by binding
to and thereby repressing the expression of the extracellular matrix
protein Fibronectin 1 (FN1) gene, but transactivatingcell migration
proteinprofilin1 (PFN1) gene [11].
In addition, in melanomas of both uveal and cutaneous origins a
G protein-related signaling pathway is frequently activated through
a gain-of-function mutation of either large GTPase α subunit like
GNA11 and GNAQ [12,16] or their homologous—small GTPase
like NRAS and its downstream effector BRAF [17,18]. We also find
that BRAF and GNA11 are transcriptionally upregulated in UM and
MetUM [11], suggesting that although GNA11/GNAQ or NRAS/
BRAF mutations are often found in uveal and cutaneous melanomas,
respectively [19], the overall elevated transcription levels of the above
genes position UM cells in a high potential readiness for extracellular
signals for cell proliferation regardless of their gain-of-function
mutation status.Finally, we downloaded two sole UM microarray
datasets (GSE22138 with 63 primary tumors and GSE44299 with
51 primary tumors) from National Center for Biotechnology
Information (NCBI) website, and divided the primary UMs in each
array into ZEB1high and ZEB1low groups based on their tumor ZEB1
expression levels to see if ZEB1 expression in primary UM can be used
to estimate metastasis. The Kaplan-Meier survival curves of these two
large cohorts show highly significant relationship between ZEB1-high
group and their metastases [11], suggesting that ZEB1 expression
levels can be used to predict UM patient prognosis in the clinic.
As summarized in Figure 1, we have recently demonstrated
that expression of ZEB1 is positively correlated with UM malignant
progression as with carcinomas, but not through EMT, which differs
from carcinomas, and that its oncogenic function is realized via direct
binding and repressing major tumor repressors like CDKN1A, ID2
and BAP1 in enhancing tumor cell dedifferentiation, proliferation,
invasion and metastasis.
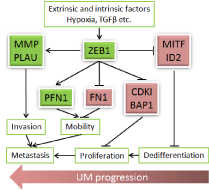
Figure 1
Figure 1
Schematic summary of ZEB1 regulation on UM progression. ZEB1
is activated by extrinsic factors like hypoxia20 and intrinsic factors like TGFβ13,
and can serve either repressor or activator depending on its co-fector13 to
directly bind and regulate expression of critical genes involved in UM
transformation, progression, and metastasis. Arrows indicate transactivation
while bars indicate repression. Red color indicatesupregulation while green
color indicates downregulation.
Acknowledgments
This work was supported by grants from theNational Natural Science Foundation of China (Grant Numbers: No. 3087282 and No. 81072221)and the Natural Science Foundation of Hunan Province (14JJ2005) and by the Basic Research Grant of University of Louisville School of Medicine (E0819) and Research to Prevent Blindness.
References
- Singh AD, Topham A. Incidence of uveal melanoma in the United States: 1973-1997. Ophthalmology. 2003;110(5):956-61.
- Yonekawa Y, Kim IK. Epidemiology and management of uveal melanoma. Hematol Oncol Clin North Am. 2012;26(6):1169-84.
- Shildkrot Y, Thomas F, Al-Hariri A, Fry CL, Haik BG, Wilson MW. Socioeconomic factors and diagnosis of uveal melanoma in the mid-southern United States. Current eye research 2011, 36(9):824-30.
- Augsburger JJ, Corrêa ZM, Shaikh AH. Effectiveness of treatments for metastatic uveal melanoma. Am J Ophthalmol. 2009;148(1):119-27.
- Blum ES, Yang J, Komatsubara KM, Carvajal RD. Clinical Management of Uveal and Conjunctival Melanoma. Oncology (Williston Park). 2016;30(1):29-32, 34-43, 48.
- Intraocular (Uveal) Melanoma Treatment (PDQ(R)): Health Professional Version. In: PDQ Cancer Information Summaries. Bethesda (MD); 2002.
- Chang SH, Worley LA, Onken MD, Harbour JW. Prognostic biomarkers in uveal melanoma: evidence for a stem cell-like phenotype associated with metastasis. Melanoma Res. 2008;18(3):191-200.
- Chou YS, Yang M. Epithelial-mesenchymal transition-related factors in solid tumor and hematological malignancy. J Chin Med Assoc. JCMA. 2015; 78(8):438-45.
- Wang Y, Zhou BP. Epithelial-mesenchymal transition in breast cancer progression and metastasis. Chin J Cancer. 2011;30(9):603-11.
- Aoyagi K, Tamaoki M, Nishumura T, Sasaki H. Technical considerations for analyzing EMT-MET data from surgical samples. Cancer Lett. 2013;341(1):105-10.
- Chen Y, Lu X, Montoya-Durango DE, Liu YH, Dean KC, Darling DS, et al. ZEB1 Regulates Multiple Oncogenic Components Involved in Uveal Melanoma Progression. Sci Rep. 2017 7(1):45.
- Asnaghi L, Gezgin G, Tripathy A, Handa JT, Merbs SL, van der Velden PA, et al. EMT-associated factors promote invasive properties of uveal melanoma cells. Molecular vision 2015, 21:919-29.
- Sánchez-Tilló E, Liu Y, de Barrios O, Siles L, Fanlo L, Cuatrecasas M, et al. EMT-activating transcription factors in cancer: beyond EMT and tumor invasiveness. Cell Mol Life Sci. 2012;69(20):3429-56.
- Harbour JW, Onken MD, Roberson ED, Duan S, Cao L, Worley LA, et al. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science. 2010;330(6009):1410-3.
- Onken MD, Ehlers JP, Worley LA, Makita J, Yokota Y, Harbour JW. Functional gene expression analysis uncovers phenotypic switch in aggressive uveal melanomas. Cancer Res. 2006;66(9):4602-9.
- Mimeault M, Batra SK. Molecular biomarkers of cancer stem/progenitor cells associated with progression, metastases, and treatment resistance of aggressive cancers. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology. 2014; 23(2):234-54.
- Yousef YA, Alkilany M. Characterization, treatment, and outcome of uveal melanoma in the first two years of life. Hematol Oncol Stem Cell Ther. 2015;8(1):1-5.
- Singh AD, Turell ME, Topham AK. Uveal melanoma: trends in incidence, treatment, and survival. Ophthalmology. 2011;118(9):1881-5.
- Duan F, Lin M, Li C, Ding X, Qian G, Zhang H, et al. Effects of inhibition of hedgehog signaling on cell growth and migration of uveal melanoma cells. Cancer biology & therapy 2014, 15(5):544-59.
- Liu Y, Clem B, Zuba-Surma EK, El-Naggar S, Telang S, Jenson AB, et al. Mouse fibroblasts lacking RB1 function form spheres and undergo reprogramming to a cancer stem cell phenotype.Cell Stem Cell. 2009, 4(4):336-47.