Research Article
Gene Expression Profiling of Colorectal Cancer by Correlation with 18F-FDG Kinetics as Measured by Dynamic Positron Emission Tomography-Computed Tomography (dPET-CT): Dependency on Cadherin-Related Genes and Hypoxia
Caixia Cheng*, Sven Klippel, Dirk Koczan, Stefan Willis, Leyun Pan, Christos Sachpekidis and Antonia Dimitrakopoulou-Strauss
Department of Nuclear Medicine, German Cancer Research Center, Germany
*Corresponding author: Caixia Cheng, Department of Nuclear Medicine, German Cancer Research Center, Heidelberg, Germany
Published: 05 Jan, 2017
Cite this article as: Cheng C, Klippel S, Koczan D, Willis
S, Pan L, Sachpekidis C, et al. Gene
Expression Profiling of Colorectal
Cancer by Correlation with 18F-FDG
Kinetics as Measured by Dynamic
Positron Emission Tomography-
Computed Tomography (dPET-CT):
Dependency on Cadherin-Related
Genes and Hypoxia. Clin Oncol. 2017;
2: 1179.
Abstract
Purpose: The kinetics of 18F-FDG as measured by dPET-CT is determined by glucose transporters
and hexokinases, which may be regulated by other genes. The dependency of 18F-FDG kinetics on
hypoxia- and cadherin-related gene expression was assessed in this study.
Procedures: Patients with colorectal tumors (n = 18) were studied with 18F-FDG dPET-CT. Tissue
specimens were obtained from the tumor and the normal colon during the process of surgery,
and then gene expression was examined using gene arrays. The dynamic PET data were assessed
using compartmental (a two-tissue compartment model) and non-compartmental models (fractal
dimension).
Results: Overall, 13 hypoxia- and cadherin-related genes were identified with a tumor-to-normal
ratio exceeding 2.0. The number of significant correlation was different for each PET parameter
using a significance level of p <0.05. Statistical analysis revealed a significant correlation between
K1 and H-cadherin 13 (CDH13) (r = 0.92) as well as between the Fractal Dimension (FD) and
Protocadherin 43 (PCDHGC3) (r = 0.63). Furthermore, we detected a significant correlation
between FD and Protocadherin gamma subfamily B, 7 (PCDHGB7) (r = 0.60), as well as between
K1 and protocadherin 17 (PCDH17) (r = 0.55). SUV was correlated with protocadherin beta 17
(PCDHB17) with r = 0.56. A correlation coefficient of r = 0.42 was found for K1 and the expression
of hypoxia-inducible protein 2 (HIG2).
Conclusion: The transport rate for 18F-FDG (K1) is higher in tumors with a high expression of
cadherin- and hypoxia-related genes. The parameters of 18F-FDG dPET kinetics may be used to
predict the expression of hypoxia and cadherin-related genes individually.
Keywords: Gene expression; DPET-CT; Colorectal tumor; Hypoxia; Cadherin
Introduction
Colorectal Cancer (CRC) is the third most common cancer, and it is also the second leading
cause of death due to cancer [1]. Current prognostic models based on histoclinical parameters are not
accurate enough to predict individual patient because the histoclinical parameters apprehend only
poorly the heterogeneity of disease. Tumors with diverse genetic alterations, developing in varied
host backgrounds, could have the same clinical presentation but follow very different evolutions.
DNA microarray technology can be used to measure the mRNA expression level of thousands of
genes simultaneously in a single assay [2]. Gene expression profiling can disclose biologically and/or
clinically relevant subgroups of tumors. So far normal and tumor samples of CRC or varied stages of
disease have been compared by profiling gene expression in many research groups.
2-Deoxy-2-18F-fluoro-D-glucose (FDG) is well known as a radiopharmaceutical of tumor
viability and is used in the Positron Emission Tomography (PET). FDG uptake in dynamic
images is dependent on several factors, covering the fractional Blood Volume (VB) of a tumor, the
glucose transport, and the phosphorylation of the intracellular FDG. The impact of these factors is dependent on the tumor histology and can be highly different.
Current PET-CT technology makes it possible to evaluate glucose
consumption quantitatively by using dynamic data acquisition
protocols and applying dedicated software programs, which is
very useful, for example, in achieving a more accurate diagnosis in
different tumor entities, like in soft tissue sarcomas [3]. Furthermore,
the kinetic FDG data obtained by applying a two-tissue compartment
model to the dPET data can be compared with gene expression data,
if tumor samples are obtained from the same region [4-5]. Pugachev
et al. [6] assessed the dependency of the FDG uptake from the tumor
microenvironment in an experimental study with prostate tumors.
Their results demonstrated that FDG is indicative for tumor hypoxia,
but neither blood flow nor cellular proliferation. In the study of Airley
and Mobasheri, the aspects of hypoxic regulation of glucose transport,
metabolism, and angiogenesis were evaluated and the link between
hypoxia, angiogenesis and glucose transporters was identified [7].
Recently, we concluded that the FDG kinetics in colorectal tumors
was modulated by angiogenesis-related gene expression in our study
[5], and the impact of cell-proliferation-associated gene expression
on FDG kinetics was also identified [8].
However, it is well known that cell-cell adhesion receptors,
like E-cadherin, play an important role by determining tumor
progression of colorectal cancer, serving as a suppressor of invasion
and metastasis. Jeanes et al. [9] suggested three potential underlying
mechanisms as an explanation of the tumor progression induced
by the loss of E-cadherin, The one mechanism is the capacity of
E-cadherin to modulate β-catenin signaling in the canonical Wnt
pathway. The second oneis its potential to inhibit mitogenic signaling
through growth factor receptors and the possible links between
cadherins and the molecular determinants of epithelial polarity
[10]. To our knowledge no data exist about the cadherin expression
in colorectal tumors and the FDG-uptake. We focus on this aspect,
because cadherins play an important role in the progression of
colorectal cancers. We know, that the FDG-uptake is regulated
by many different molecular mechanisms, like proliferation and
angiogenesis. The topic of this paper is to evaluate the correlation
between cell adhesion molecules and the FDG-uptake, which is an
unspecific feature present in different tumors including colorectal
cancer.
Here we have applied DNA microarray technology to analyze
the expression of 54,675 genes in 18 cancerous (primary tumors) and
noncancerous colon tissue samples (reference tissue), and to assess
the impact of hypoxia and cadherin-associated genes on the FDG
kinetics in primary colorectal tumors.
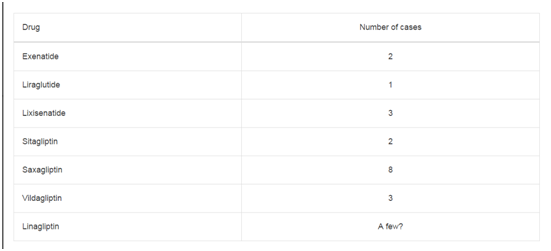
Table 1
Materials and Methods
Patient data
A total of 18 patients including 10 men and 8 women with an
average age of 65 as shown in Table 1 have been diagnosed with
histologically confirmed colorectal tumors. PET-CT studies with
18F-FDG were performed in these patients before surgery. Only
patients with tumors exceeding 3 cm were considered for the data
evaluation in this study. All PET-CT studies were performed within
2 days before surgery.
Patients providedus with written informed consent to participate
in the study and to have their medical records released. The Ethical
Committee I of the University of Heidelberg and the Federal Agency
for Radiation Protection (Bundesamt für Strahlenschutz) approved
the research project.
Dynamic PET/CT studies
Dynamic PET/CT examinations with 18F-FDGwere routinely
performed over the area of the knowntumor for each patient
using a 28-frame protocol for 60 min at our center. We injected
intravenously a maximum of 250 MBq18F-FDG. The dynamic data
acquisition protocol covers 10 frames of 30 seconds, 5 frames of 60
seconds, 5 frames of 120 seconds and 8 frames of 300 seconds, which
is increasing time per frame. After the end of the dynamic study we
acquired additional static images from the maxilla to the knees based
on the movement of the table in the craniad and caudad directions.
More details about the procedure and PET/CT system used in this
study are already described [5]. All PET images were attenuationcorrected
and an image matrix of 400 x 400 pixels was used for
iterative image reconstruction. Iterative images reconstruction was
based on the Ordered Subset Expectation Maximization Algorithm
(OSEM) with six iterations and twelve subsets. The reconstructed
images were converted to SUV images based on the formula [11]:
SUV = tissue concentration (Bq/g)/(injected dose (Bq)/body weight
(g)).
Data acquisition and data evaluation
The quantitative evaluation of the dynamic PET data was
performed with dedicated software PMOD (PMOD Technologies
Ltd, Zürich, Switzerland), which was developed from our project
group. The process of the PET data acquisition using PMOD
software was described elsewhere [8]. The quantitative evaluation
of the dynamic PET studies was performed by fitting a 2-tissuecompartment
model to the VOI data, which uses an input function
to provide the 18F-FDG concentration in the vessels. Ohtake et al.
[12] displayed that the input data for 18F-FDG could be accurately
obtained via ROIs over large blood vessels [12]. Therefore, we placed
at least 7 consecutive ROIs over the descending aorta to obtain the
blood data for 18F-FDG. The recovery coefficient is 0.85 for a diameter of 8 mm using our reconstruction protocol. Because the diameter of
all VOIs exceeded 8 mm, no partial-volume correction was applied.
After the placement of VOIs for the tumor, the normal colon, and
the blood vessels, the 2-tissue model was fitted iteratively to obtain
the compartment parameters. The compartmental model provided us
with five parameters of K1-k4 and VB. The rate constants (K1-k4)
have the unit 1/min, whereas VB is associated with the fraction of
blood within the evaluated target volume. The constants K1 and k2
are associated with 18F-FDG transport; the constants k3 and k4 are
correlated with the phosphorylation and de phosphorylation of the
intracellular 18F-FDG; and the parameter VB reflected the fractional
blood volume in the target volume, which also referred to as vessel
density. The measured data were fitted by minimizing the summed
square differences between estimated and measured values. Results
were commonly accepted as valid if K1–k4 were less than 1 and VB exceeded 0. The global influx of 18F-FDG was calculated from
the compartment data using the following equation: influx (INF) =
(K1 · k3)/(k2 + k3).In addition, the Fractal Dimension (FD) of the
time–activity curve was calculated using a no compartment model,
which varies from 0 to 2, and provides information about the more
deterministic or chaotic distribution of the tracer over time.
Tissue specimen and gene arrays
The tissue specimens of the tumor and normal colon were
transported in liquid nitrogen, and total RNA was extracted for
further processing. The quality of isolated RNA was assessed photo
metrically using the 280:260 ratio and on an agarose gel as described
before [8]. We used the U133 Plus 2.0 gene chip (Affymetrix Inc.),
which provides quantitative information about 54,675 gene probes.
The processing of the RNA and gene arrays was performed according
to the manufacturer’s recommendations as described elsewhere [13].
Gene chip–expression data were normalized for the β2-microglobulin
(Affymetrix code 34644_at, Homo sapiens mRNA for β2-
microglobulin) using the following equation [4]: relative expression
value (REV) = 1,000 · (expression value of a gene/expression value for
β2-microglobulin).
Statistical data evaluation
The statistical evaluation was performed with Stata/SE 10.0 (Stata
Corp.) on a Mac Pro (Apple Inc.) 2 · 3 GHz Quad-Core Xeon system
(Intel Corp.) with 16 GB of RAM, using Mac OS X 10.5.1. The same
system was also used for all data-processing tasks in this study. We
developed dedicated software GenePET for the correlative evaluation
of dynamic PET and gene-array data [13], which can store both genearray
and PET data as well as the individual gene codes. Meanwhile, a
full description of each gene and its function is also stored.
Table 2
Table 2
Max., Min., Mean, Median and SD of Patients for Tumors and Reference
Tissue (Normal Colon) for PET Results (The number of patients: n=18).
Figure 1
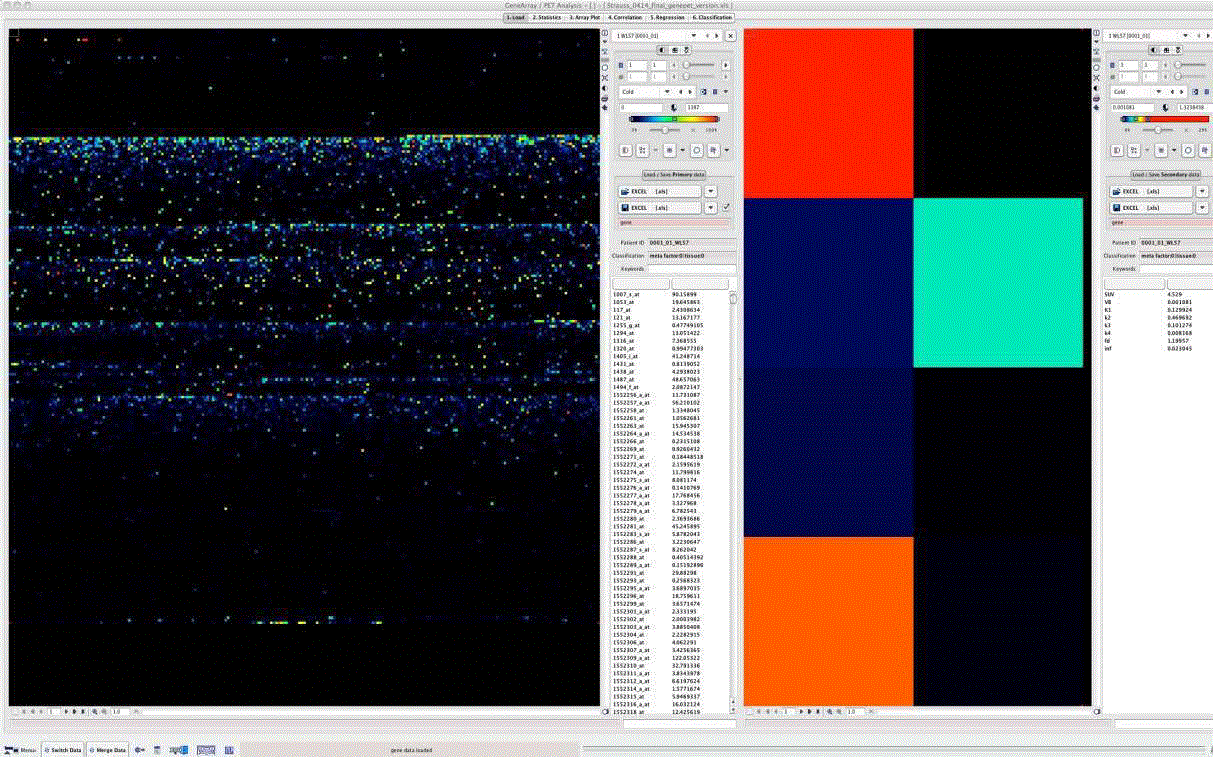
Figure 1
GenePET display of data obtained with gene array from tumor sample (left) and corresponding quantitative kinetic PET data (right). Both datasets are
displayed as an image to facilitate detection of enhanced genes.
Figure 2
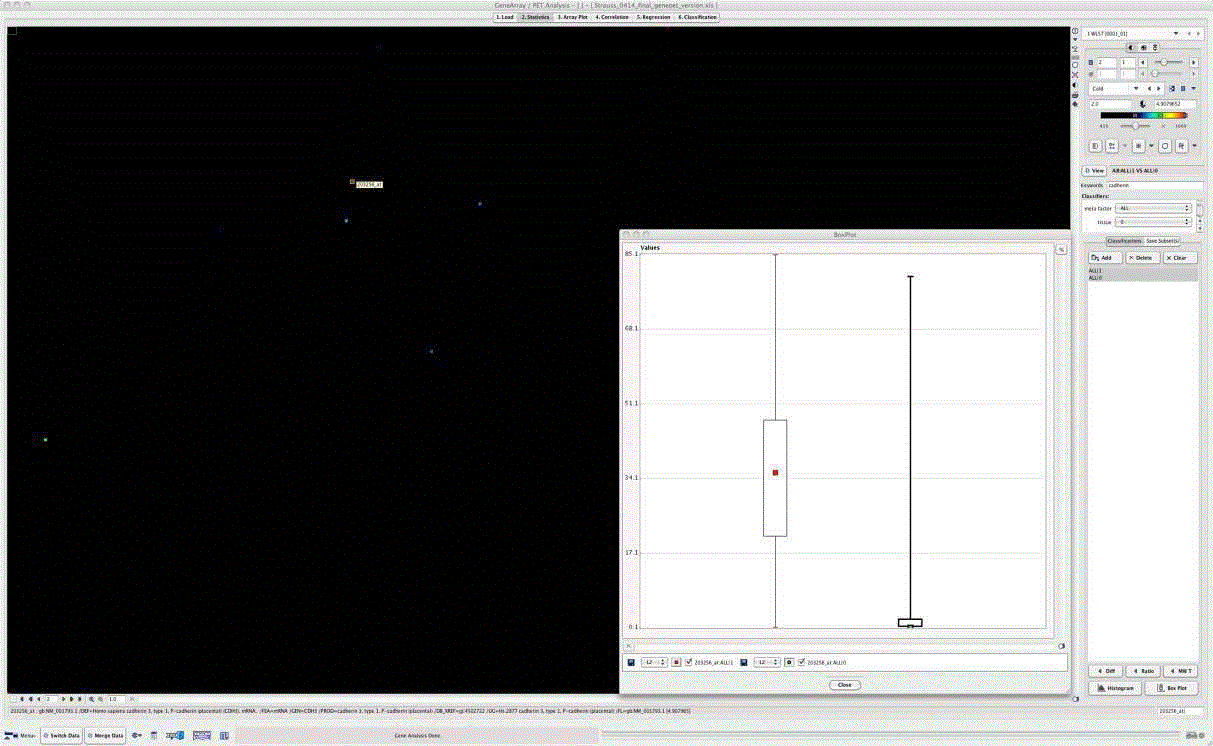
Figure 2
Tumor-to-normal tissue rations for genes related to cadherin. Ratios are calculated from pairwise gene-expression data for tumor and normal colon of
each patient. Key words cadherin and hypoxia were used for selection and revealed 13 associated genes. Highest ratio (4.91) was found for P-cadherin (CDH3)
(gene code 203256_at).
Results
Overall, tissue specimens of the tumor and the normal colon
were obtained in 18 patients. Therefore, we were able to obtain geneexpression
data and dPET-CT studies in 18 patients. The basic results
for PET parameters are shown in Table 2.
The Gene PET software we developed was used to identify
differentially enhanced genes related to the hypoxia and cadherin
(Figure 1 and 2). All probe values were stored together with the
Affymetrix code and a full description of each probe in a dedicated
format. First, tumor-to-colon ratios were calculated by using the
tumor and normal samples of each patient. We calculated the
tumor-to-colon tissue ratio based on patient-by-patient, by using
the corresponding gene-expression values of the tumor and the
reference tissue data (from normal colon mucosa) of the same
patient. Furthermore, the keyword search with “hypoxia or cadherin”
according to the stored description of the genes was used to select
only the hypoxia- and cadherin-related genes from the list of all
probes (Figure 2). Overall the search algorithm revealed 159hypoxiaand
cadherin-related genes. Thehypoxia- and cadherin-related genes
with tumor-to-colon ratios (median values) exceeding two are given
in Table 3. 13 hypoxia and cadherin-related genes revealed a tumorto-
colon ratio >2. The highest tumor-to-colon ratio was obtained
from P-cadherin (CDH3) with a value of 4.91.
We used the median value for K1 (K1 = 0.23) to classify
the tumor data into 2 groups with low and high K1 values. The
classification analysis revealed that low K1 values were associated
with low expression of CDH13 using the Wilcoxon rank sum test.
The results were highly statistically significant (p = 0.0006) (Figure
3). Meanwhile, correlation analysis was used for the tumor specimen
to identify dependencies of the 18F-FDG kinetics on hypoxia- and
cadherin-related gene expression. The pair wise correlation using
a significance level of p <0.05 was applied to the 13 selected genes
by key words “hypoxia or cadherin”. The results demonstrated that
the number of significant correlations was different for each PET
parameter; for example, 8 significant correlations were available
for K1. Only the most significant correlations were used for further
regression analysis. The analysis revealed the highest correlation
between K1 and H-cadherin 13 (CDH13) with r = 0.92 (p <0.001).
Furthermore, we found correlation between Fractal Dimension (FD)
and Protocadherin 43 (PCDHGC3) with r = 0.63 (p <0.01), as well as
between FD and Protocadherin gamma subfamily B, 7 (PCDHGB7)
with r=0.60(p< 0.01). A significant correlation was also demonstrated
between K1 and protocadherin 17 (PCDH17) with r = 0.55 (p <0.02).
SUV was correlated with protocadherin beta 17 (PCDHB17) with
r = 0.56 (p <0.02). A correlation coefficient of r = 01.42 (p <0.05)
was noted for K1 and the expression of hypoxia-inducible protein 2
(HIG2). All results were listed in the last column of Table 3.
Discussion
Gene arrays are very useful in evaluating the expression of a
large group of genes. The Gene PET software makes it possible to
select subgroups of genes using a key word search according to the
description of the genes. The kinetics of FDG is primarily displaying
the transport and phosphorylation of glucose. Therefore, the
evaluation of FDG kinetics using a 2-tissue compartment model can
not only help us to quantify effects but also provide us data about
molecular biological aspects. However, FDG kinetics is regulated by
many other mechanisms, like angiogenesis and proliferation. Riedl et
al. [14] evaluated the FDG uptake in patients with metastatic colorectal
cancer and found that FDG provides predictive information in these patients. So far there is no information reflecting the correlation of the
FDG uptake and markers of cell adhesion and hypoxia. When a tumor
grows, it rapidly outgrows its blood supply, leaving portions of the
tumor with regions where the oxygen concentration is significantly
lower than in healthy tissues. In order to support continuous
growth and proliferation in challenging hypoxic environments,
cancer cells are detected to alter their metabolism [15]. Expression
of genes related to glycolytic enzymes and glucose transporters are
enhanced by numerous oncogenes including RAS, SRC, and MYC
[16,17]. Commonly, hypoxia gives rise to increase the production
of hypoxia-inducible factors (HIF-1), including HIF-1α and HIF-1β
subunits, which act as a key regulatory transcription factor related
to adaptive cellular changes. In humans, HIF-1 has been verified to
up-regulate expression of genes affecting a range of target areas of
physiology. These genes include not only those involved in triggering
an inflammatory response but also those related to iron metabolism.
It is especially notable that HIF-1 is shown to affect glycolytic genes
to interfere with reductions in oxygen availability and consumption
when focusing on metabolism. GLUT-1, HIF-1α and the Proliferating
Cell Nuclear Antigen (PCNA) in 60 tumor specimens of colorectal
tumors and 20 normal colon probes were evaluated by Zhou and
Deng [18]. Their results demonstrated an over expression of both
GLUT-1 and HIF-1α as well as a close correlation of the genes with
cellular proliferation as measured by PCNA. The experimental
studies from Burgman et al. [19] on MCF7 cells also demonstrated
an increase of 18F-FDG uptake induced by hypoxia. The effect of
VEGF and GLUT-3 on hypoxia-inducible factor-1 was modulating
by both VEGF and GLUT-3 (20). The results disclosed hierarchical
dependencies of glucose transporter expression on angiogenesis
and hypoxia. Pedersen et al. [20] found a significant up-regulation
of VEGF, GLUT-1 and GLUT-3 during hypoxia when they used 2
human small-cell lung cancer cell lines [21]. Similarly, a significant
correlation of HIF-1α and GLUT-1 expression in patients with renal
cell carcinomas was also found by Lidgren et al. [22]. In addition, the
effect of hypoxia on the deoxyglucose uptake and cell cycle regulatory
protein expression of mouse embryonic stem cells were evaluated by
Lee et al. [23]. Under hypoxia the authors verified a co-up-regulation
of cell cycle regulatory protein expression, including cdk2 and GLUT-
1.In this study, we found a low but significant correlation betweenK1
and the expression of hypoxia-inducible protein 2 (HIG2).The results
demonstrate that there is a dependency between hypoxia and FDGuptake.
However, more data are needed to verify it.
The combination of Gene PET and dynamic PET parameters is a
more sensitive and accurate method to obtain information about the
metabolism of a tumor, which is modulated by different parameters
including hypoxia. We also found an increased expression of hypoxiainducible
protein 2 (HIG2). Furthermore, we found a significant
correlation for K1 of a two-tissue compartment model, which reflects
the transport of 18F-FDG into the cells, and the expression of hypoxiainducible
protein 2 (HIG2) as measured by a gene-array techniques.
However, no significant over expression of HIF-1α (Tumor-to-colon
Ratio = 1.14) was found.
Cadherins are a category of type-1transmembrane proteins and
play important roles in cell adhesion and recognition. Cadherins
behave as both receptors and ligands for other molecules. During
the developing process, they behave as an assist in properly
positioning cells: they are related to the separation of the different
tissue layers, and responsible for cellular migration [24]. E-cadherin
(epithelial cadherin) is most greatly expressed in the very early
stages of development. N-cadherin (neural cadherin) is expressed
and E-cadherins down-regulate expressed during the next stage of
the development of the neural plate. Finally, E- P- and N-cadherin
expression increases during the development of the notochord and the condensation of somites. Cadherins play a role in maintaining
cell and tissue structure as well as in cellular movement after the end
of development [25]. In addition, regulation of cadherin expression
can occur bypromoting methylation among other epigenetic
mechanisms [26]. In this study, the highest tumor-to-normal tissue
ratio, 4.91, was found for P-cadherin (CDH3). In addition, we also
found a highly significant correlation for K1 and the H-cadherin
(CDH13) expression (Figure 3). The loss of H-cadherin in tumor cells
is correlated with tumor malignancy, invasiveness and metastasis.
Toyooka S et al. [27] reported that tumor progression in colorectal
cancer associates with down regulation of Т-cadherin expression,
which is consistent with our results. Wang XB et al. [28] found
Protocadherin-17(PCDH17) promote methylation, which is closely
associated with bladder cancer malignancy. In this study, our results
revealed a significant correlation between K1 and PCDH17, which is
the first report in colorectal cancer. Protocadherin 43 (PCDHGC3)
and Protocadherin gamma subfamily B, 7 (PCDHGB7) are both
the member of the protocadherin gamma gene cluster. These gene
clusters have an immunoglobulin-like organization, displaying
that a novel mechanism may be involved in their regulation and
expression [29]. However, their exact role is poorly understood.
We noted a significant correlation between Fractal Dimension (FD)
and Protocadherin 43 (PCDHGC3) (r = 0.63) as well as FD and
Protocadherin gamma subfamily B, 7 (PCDHGB7) (r = 0.60). The
FD was calculated from the dynamic PET data. It reflects the more
chaotic or deterministic distribution of the 18F-FDG kinetics, which
may explain the association with protocadherin family, which plays
a very important role in the establishment and function of specific
cell-cell connections.
Table 3
Table 3
Tumor-to-colon Ratios exceeding 2 for Hypoxia- and Cadherin-Related Genes identified with GenePET program using key words.
Figure 3
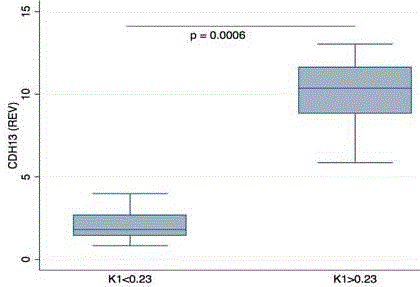
Figure 3
Box plot of K1 and the expression of CDH13 for tumor specimen.
Median of K1 was used to classify data. Groups were significantly different
using Wilcoxon rank sum test (p=0.0006). REV: relative expression value.
Conclusion
Our results verify that cadherin is a worth studying parameter for the 18F-FDG kinetics in colorectal tumors. The correlation between the PET parameters and the gene-expression data makes it possible to noninvasively predict the expression of these cadherin-related genes in patients with primary colorectal carcinomas based on the dynamic PET data. Further studies are needed to evaluate the quantitative impact of these biologic parameters on the 18F-FDG kinetics.
References
- Kosmidou V, Oikonomou E, Vlassi M, Avlonitis S, Katseli A, Tsipras I, et al. Tumor heterogeneity revealed by KRAS, BRAF, and PIK3CA pyrosequencing: KRAS and PIK3CA intratumor mutation profile differences and their therapeutic implications. Human Mutation. 2014; 35: 329-340.
- Bertucci F, Salas S, Eysteries S, Nasser V, Finetti P, Ginestier C, et al. Gene expression profiling of colon cancer by DNA microarrays and correlation with histoclinical parameters. Oncogene. 2004; 24: 1377-1391.
- Dimitrakopoulou-Strauss A, Strauss LG, Schwarzbach M, Burger C, Heichel T, Willeke F, et al. Dynamic PET 18F-FDG studies in patients with primary and recurrent soft-tissue sarcomas: impact on diagnosis and correlation with grading. J Nucl Med. 2001; 42: 713-720.
- Strauss LG, Dimitrakopoulou-Strauss A, Koczan D, Bernd L, Haberkorn U, Ewerbeck V, et al. 18F-FDG kinetics and gene expression in giant cell tumors. J Nucl Med. 2004; 45: 1528-1535.
- Strauss LG, Koczan D, Klippel S. Impact of angiogenesis-related gene expression on the tracer kinetics of 18F-FDG kinetics in colorectal tumours. J Nucl Med. 2008; 49: 1238-1244.
- Pugachev A, Ruan S, Carlin S, Larson SM, Campa J, Ling CC, et al. Dependence of FDG uptake on tumour microenvironment. Int J RadiatOncolBiol Phys. 2005; 62: 545-553.
- Airley RE, Mobasheri A. Hypoxic regulation of glucose transport, metabolism and angiogenesis in cancer: novel pathways and targets for anticancer therapeutics. Chemotherapy. 2007; 53: 233-256.
- Strauss LG, Koczan D, Klippel S. Impact of cell-proliferation-associated gene expression on 2-Deoxy-2-18Fluoro-D-glucose (FDG) kinetics as measured by dynamic positron emission tomography (dPET) in colorectal tumors. Mol Imaging Biol. 2011; 13: 1290-1300.
- Jeanes A, Gottardi CJ, Yap AS. Cadherins and cancer: how does cadherin dysfunction promote tumor progression? Oncogene. 2008; 27: 6920-6929.
- Qian X, Karpova T, Sheppard AM, McNally J, Lowy DR. E-cadherin-mediated adhesion inhibits ligand-dependent activation of diverse receptor tyrosine kinase. EMBO J. 2004; 23: 1739-1784.
- Straus LG, Conti PS. The application of PET in clinical oncology. J Nucl Med. 1991;1 32: 623-648.
- Ohtake T, Kosaka N, Watanabe T, Yokoyama I, Moritan T, Masuo M, et al. Noninvasive method to obtain input function for measuring glucose utilization of thoracic and abdominal organs. J Nucl Med. 1991; 32: 1432-1438.
- Strauss LG, Pan L, koczan D. Fusion of Positron Emission Tomography (PET) and gene array data: a new approach for the correlative analysis of molecular biological and clinical data. IEEE Trans Med Imaging. 2007; 26: 804.
- Riedl CC, Akhurst T, Larson S. 18F-FDG PET scanning correlates with tissue markers of poor prognosis and predicts mortality for patients after liver resection for colorectal metastases. J Nucl Med. 2007; 48: 771-775.
- Vander H, Matthew G, Lewis CC, Craig BT. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009; 324: 1029-1033.
- Flier JS. Elevated levels of glucose transport and transporter messenger RNA are induced by ras or src oncogenes. Science. 1987; 235: 1492-1495.
- Osthus RC. Deregulation of glucose transporter 1 and glycolytic gene expression by c-Myc. J Biol. Chem. 2000; 275: 21797-21800.
- Zhou YL, Deng CS. Correlations of expression of Glut1 and HIF-1alpha to cellular proliferation of colorectal adenocarcinoma. Ai Zheng. 2005; 24: 685-689.
- Burgman P, O’Donoghue JA, Humm JL, Ling CC. Hypoxia-induced increase in FDG uptake in MCF7 cells. J Nucl Med. 2001; 42:170-175.
- Maxwell PH, Dachs GU, Gleadle JM, Nicholls LG, Harris AL, Stratford IJ, et al. Hypoxia-inducible factor-1 modulates gene expression in solid tumors and influences both angiogenesis and tumor growth. ProcNatlAcadSci USA. 1997; 94: 8104-8109.
- Pedersen MW, Holm S, Lund EL. Coregulation of glucose uptake and vascular endothelial growth factor (VEGF) in two small cell lung cancer (SCLC) sublines in vivo and in vitro. Neoplasia. 2001 3: 80-87.
- Lidgren A, Bergh A, Grankvist K, Rasmuson T, Ljungberg B. Glucose transporter-1 expression in renal cell carcinoma and its correlation with hypoxia inducible factor-1a. BJU int. 2008; 101: 480-484.
- Lee SH, Heo JS, Han HJ. Effect of hypoxia on 2-deoxyglucose uptake and cell cycle regulatory protein expression of mouse embryonic stem cells: involvement of Ca2+/PKC, MAPKs and HIF-1alpha. Cell PhysiolBiochem. 2007; 19: 269-282.
- Gumbiner BM. Regulation of cadherin-mediated adhesion in morphogenesis. Nature Reviews Molecular cell Biology. 2005; 622-634.
- Tepass U, Truong K, Godt D, Ikura M, Peifer M. Cadherins embryonic and neural morphogenesis. Nature Reviews Molecular cell Biology. 2000.
- Reinhold WC, Reimers MA, Maunakea AK, Kim S, Lababidi S, Scherf U, et al. Detailed DNA methylation profiles of the E-cadherin promoter in the NCI-60 cancer cells. Molecular cancer therapeutic. 2007; 6: 391-403.
- Toyooka S, Toyooka KO, Harada K, Miyajima K, Makarla P, Sathyanarayana UG, et al. Aberrant methylation of the CDH13 (H-cadherin) promoter region in colorectal cancers and adenomas. Cancer Res. 2002; 62: 3382-3386.
- Wang XB, Lin YL, Li ZG. Protocadherin 17 promoter methylation in tumor tissue from patients with bladder transitional cell carcinoma. J Int Med Res. 2014; 42: 292-299.
- Wu Q, Maniatis T. A striking organization of a large family of human neural cadherin-like cell adhesion genes. Cell. 1999; 97: 779-790.