Commentary
Birth Order in Malignant Hematological Disorders: A Challenge
Viggo Jønsson*
Department of Hematology, Rikshospital and Institute of Clinical Medicine, University of Oslo, Norway
*Corresponding author: Viggo Jønsson, Department of
Hematology, Rikshospital, Institute of
Clinical Medicine, University of Oslo,
Norway
Published: 01 Jul, 2016
Cite this article as: Jønsson V. Birth Order in Malignant
Hematological Disorders: A Challenge.
Clin Oncol. 2016; 1: 1053.
Commentary
A birth order effect (BOE) is a non-random occurrence of affected siblings in a sib ship. Thus, in case of BOE the affected child has a specific and significant birth rank order by age, for example first child, early in the sib ship, in the middle or late [1].
Occurrence
Birth order was originally discovered in families with congenic malformations. Sewall Wright was one of the pioneers within this field around the time nineteen twenties. He was well aware of the fact that BOE is not in accordance with a traditional Mendelian segregation. He also described a relationship between parents age and BOE, in polydactyl for example with a maternal age effect, and outlined the empiric risk for genetic counselling based on the general assumption that the presence of the mutation, sporadic versus inherited, which causes the malformation, could be related to parents age [1]. It is obvious that the study of BOE in a genetic disorder with unknown mode of transgenerational passage may contribute to the pathophysiological understanding, simply addressing the question “What is the genetic segregation in this inherited disease?” The malignant hematological disorders (MHD), both the lymphoproliferative disorders (LPD) and myeloproliferative disorders (MPD) are evidently genetic disorders in the sense that the susceptibility to disease is inherited. Thus, genome-wide-screenings have provided substantial evidence for the presence of susceptibility risk genes in a number of the diagnoses within MHD, especially in chronic lymphocytic leukemia, CLL [2-8], acute lymphoblastic leukemia (All) [9,10], monoclonal gammopathy of uncertain significance, MGUS [11,12] and in multiple myeloma [13,14]. Such diseases cannot develop unless inherited susceptibility is present in the genome. Whether persons with inborn susceptibility to MHD will develop disease or, alternatively, remain silent carriers is far from fully understood and a delicate matter for further investigation. Epigenetic and exogenous stimuli such as lymphotrophic infections seem to be factors of possible relevance [15-17]. The susceptibility genes represent the etiology to each of the diagnoses in MHD which pave the way for first step in the pathogenesis, namely a specific mutation to each of the diagnoses in either the lymphoproliferative- or the myeloproliferative differential pathway, where all diagnoses have a common characteristic: A mutated monoclone, expanding with autonomic proliferation outside the growth regulation otherwise seen in the production of normal blood cell.
Segregation and Birth Order Effectthods
Susceptibility to several diagnoses within MHD can be present in the same family. This so called pleiotropic pattern is clearly seen in genealogical investigations of pedigrees to affected families without convincing signs of a Mendelian pattern in the transgenerational segregation of the susceptibility genes [18,19]. Pseudo-dominant segregation, when the inheritance of recessive genes mimics a dominant Mendelian pattern, has been discussed [20], but not definitely confirmed so far. Genealogical investigations of family trees from affected Scandinavian families, especially CLL [18,19] confirm that the predominant element in the segregation of CLL is a parent – offspring pair, in which both the parent and the offspring is affected, and that the pattern of segregation is supposed to be due to mono-allelic genes influenced by epigenetic stimuli in such a way that no Mendelian mode is detectable [21]. A possible BOE in the transgenerational transfer of MHD-susceptibility has been hotly debated with a remarkable inconsistency in published findings. In CLL, genealogical studies showed BOE while other studies based on large-scaled screenings of cohort-data from cancer registries did not (for review see 18,19). In ALL, two recent large-scaled screenings, each with a very high sample size, detected BOE [22,23] while no BOE could be seen from screening of a Danish material [24]. BOE was not seen in AML [25] and not in Hodgkin’s lymphoma [26]. Furthermore, a relationship between BOE and effect of treatment has been described, having siblings and increasing birth order were associated with reduced survival in ALL and AML [27]. BOE is undoubtedly weak and varyingly expressed in the different diagnoses within the entity of MHD. If BOE is a result of epigenetic mechanism it must be assumed that the BOE is different from population to population, and thus different in the cohorts studied because the epigenetic stimuli, including stimuli from lymphotrophic infections, vary over time and place. It makes no sense to search for BOE unless the genealogy is completely safe and each family tree included is adjusted for miscarriages, stillbirths and extramarital children.This indeed, is a great technical and practical challenge that actually requires the family and ethical committee’s permission to inquire about other family members, to verify the diagnoses in the national cancer registry, and to talk about healthy family members, alive or dead. The necessary confidential genealogical information obtained by face-to-face interview with a patient presupposes that the doctor and the patient know each other well and that all information will remain anonymous and unrecognizable.In addition, the number of cases of MHD detected will always be lower than the real number, particularly with regard to low-grad malignant disorders with few or no symptoms, for example overlooked stage A CLL and MGUS. Furthermore, family members with susceptibility but without disease will escape undetected throughout the investigation of pedigrees. Finally, it is a weakness about birth order estimation that the cohorts for study will always contain living persons who theoretically later in life may develop disease and therefore should have been included. There is a great risk of overlooking BOE and thereby detecting false negative BOE as typical Type 2 error. This risk increases when examined for BOE in today's families with a small number of children compared to families of yesteryear.
Pathogenesis
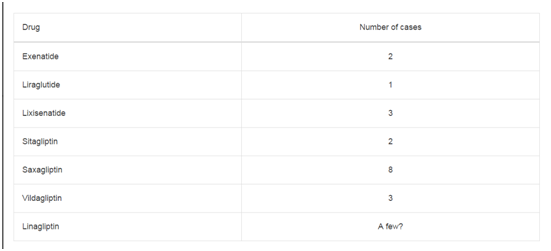
In CLL, but not in the other malignant lymphoproliferative disorder, a patrilineal but no matrilineal birth order effect has been observed among male siblings in Scandinavian families [18,19]. Is this position of male CLL patients late in the sibship in patrilineal inheritance (Table 1) explained by acquired tolerance in the mother. In this scenario, the physiological pregnancy related microchimerism between mother and fetus is caused by a two-way traffic of lymphoand monocytes between mother and fetus across the utero-placental barrier [28-32]. Thus, the mother may accumulated increased tolerance to the non-self from the paternal half of the fetus with increased number of pregnancies, births, abortions and stillborn children, and with an increased number of male partners so that eventually, the mother will be able to tolerate and to accept the paternal CLL susceptibility and the paternal susceptibility to pleiotropic diagnoses within MHD in spite of an underlying mechanism based on maternal imprinting. We now have proof that the CLL susceptibility can be accumulated. In some few of our pedigrees where both parents have CLL we see a very high occurrence of CLL among their children and we interpret this as an additive effect [19]. It is a different situation than the accumulated maternal susceptibility because of tolerance but it shows that susceptibility potentially can accumulate. Male predominance is well known in CLL and may be explained by this combination of accumulated maternal tolerance and maternal imprinting together with the presence of an allele-specific, epigenetic modifier, such as a silencer in which DNA methylation and modifications of histone are potential mechanism [21]. More than ten years ago it was shown that such specific mono-allele imprinted susceptibility genes to CLL have a higher degree of asynchronous replication compared with bi-allelic genes [33]. This asynchronous replication could be the likely supplier of the repertoire of diversity and pleiotrophy which otherwise could not be explained from a Mendelian segregation. The relationship between imprinted genes and the regulation of fetal growth factors is well established [21].There is a long and fascinating history behind this biological mechanism [34-38] which also includes what was previously called meiotic drive [39,40].Regarding the LPD-susceptibility, it could be related to the benefit of, at an early stage of fetal life, of having a defense against viral infections in terms of M-component producing lymphocytes specifically against virus, where retrovirus has been particularly under consideration [41,42]. Thus, the susceptibility genes code favorably for vital growth factors in fetal life, but later in life the susceptibility is a dangerous genetic baggage no matter the patent’s age. Combined with epigenetic stimuli, the susceptibility can cause a mono-clone of blood cells and hence a manifest malignant hematological disease. The relationship between changes in fetal growth factors and the risk of MHD could be reflected in the fact that gestational age, birth weight, increased frequency of maternal miscarriages, stillbirths and assisted reproduction are risk factors in childhood leukemia [22,43-45]. Biologically, the fetal events are undoubtedly the most important factor, giving the embryo the optimal conditions for growth, while the risk of malignant disease later on in life is hardy of any major importance since most of the malignant hematological disorders, with the exception of ALL, Hodgkin’s lymphoma and a few other cases, are mainly seen in people after their fertile life and without any influence on their reproduction. If this is so, then it is an advantage to have the dangerous genetic susceptibility uneven in the sibship in term of a BOE so that the predominant proportion of susceptibility, the patrilineal male susceptibility, is directed towards sibs who have healthy older siblings. From a genetic point of view, this means that the mother during the pregnancy directs the father-son susceptibility to sons who have unaffected older siblings. Referring to the mother-fetus two-way interaction, there can be a net epigenetic and microchimeristic traffic from first child to mother to second child to mother, etc. Unaffected in this context means sons who do not have the maternally imprinted gene or sons who have this gene as silent bystander without disease. Unfortunately, we do not know much about such unaffected family members [46]. In the other lines: father-daughter, mother-son and mother- daughter, we do not see such a BOE. The main part of the father-son susceptibility comes naturally from father-son transmissions and, theoretically, also from the mother's genome as a genetic baggage she has from affected male ancestors.If so, an accumulation of patrilineal male susceptibility takes place. It is tempting to see this as an explanation for the male predominance in CLL.
Acknowledgement
Thanks to Professor Neil D. Jones, Department of Computer Science, University of Copenhagen, Denmark, for valuable feedback to the calculations and for comments to the manuscript.
Table 1
References
- Emery AEH. Parental age and birth order. In: Emery AEH (ed.) Methodology in medical genetics, 2nd Edition. Edinburgh: Churchill Livingstone; 1986. p. 40-53.
- Gruber M, Wu CJ. Evolving understanding of the CLL genome. Semin Hematol. 2014; 51: 177-187.
- Shaknovich R. Gene expression and epigenetic deregulation. Adv Exp Med Biol. 2013; 792: 133-150.
- Slager SL, Caporaso NE, de Sanjose S, Goldin LR. Genetic susceptibility to chronic lymphocytic leukemia. Semin Hematol. 2013; 50: 296-302.
- Speedy HE, Sava G, Houlston RS. Inherited susceptibility to CLL. Adv Exp Med Biol. 2013; 792: 293-308.
- Quesada V, Ramsay AJ, Rodríguez D, Puente XS, Campo E, López-Otín C. The genomic landscape of chronic lymphocytic leukemia: clinical implications. BMC Med. 2013; 11: 124.
- Sava GP, Speedy HE, Houlston RS. Candidate gene association studies and risk of chronic lymphocytic leukemia: a systematic review and meta-analysis. Leuk Lymphoma. 2014; 55: 160-167.
- Berndt SI, Camp NJ, Skibola CF, Vijai J, Wang Z, Gu J, et al. Meta-analysis of genome-wide association studies discovers multiple loci for chronic lymphocytic leukemia. Nat Commun. 2016; 7: 10933.
- Liao F, Yin D, Zhang Y, Hou Q, Zheng Z, Yang L, et al. Association Between PIP4K2A Polymorphisms and Acute Lymphoblastic Leukemia Susceptibility. Medicine (Baltimore). 2016; 95: e3542.
- Shi Y, Du M, Fang Y, Tong N, Zhai X, Sheng X, et al. Identification of a novel susceptibility locus at 16q23.1 associated with childhood acute lymphoblastic leukemia in Han Chinese. Hum Mol Genet. 2016.
- Landgren O, Kristersson SY, Goldin LR, Caporaso NE, Blimark C, Mellquist UH, et al. Risk of plasma cell and lymphoproliferative disorders among 14621 first-degree relatives of 4458 patients with monoclonal gammopathy of undetermined significance in Sweden. Blood 2009; 114: 791-795.
- Weinhold N, Johnson DC, Rawstron AC, Försti A, Doughty C, Vijayakrishnan J, et al. Inherited genetic susceptibility to monoclonal gammopathy of unknown significance. Blood. 2014; 123: 2513-2517.
- Dimopoulos K, Gimsing P, Grønbæk K. The role of epigenetics in the biology of multiple myeloma. Blood Cancer J. 2014; 4: e207.
- Morgan GJ, Johnson DC, Weinhold N, Goldschmidt H, Landgren O, Lynch HT, et al. Inherited genetic susceptibility to multiple myeloma. Leukemia. 2014; 28: 518-524.
- McNally RJ, Eden TO. An infectious aetiology for childhood acute leukaemia: a review of the evidence. Br J Haematol. 2004; 127: 243-263.
- Marcotte EL, Ritz B, Cockburn M, Yu F, Heck JE. Exposure to infections and risk of leukemia in young children. Cancer Epidemiol Biomarkers Prev. 2014; 23: 1195-1203.
- Rudant J, Lightfoot T, Urayama KY, Petridou E, Dockerty JD, Magnani C, et al. Childhood acute lymphoblastic leukemia and indicators of early immune stimulation: a Childhood Leukemia International Consortium study. Am J Epidemiol. 2015; 181: 549-562.
- Jønsson V, Tjønnfjord G, Samuelsen SO, Johannesen T, Olsen J, Sellick G, et al. Birth order pattern in the inheritance of chronic lymphocytic leukemia and related lymphoproliferative disorders. Leuk& Lymph 2007; 48: 2387-96.
- Jønsson V, Tjønnfjord GE, Johannesen TB, Ly B, Olsen JH, Yuille M. Familial chronic lymphocytic leukemia in Norway and Denmark. Comments on pleiotropy and birth order. In Vivo. 2010; 24: 85-95.
- Yuille MR, Matutes E, Marossy A, Hilditch B, Catovsky D, Houlston RS. Familial chronic lymphocytic leukaemia: a survey and review of published studies. Br J Haematol. 2000; 109: 794-799.
- Sasaki H, Matsui Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat Rev Genet. 2008; 9: 129-140.
- Von Behren J, Spector LG, Mueller BA, Carozza SE, Chow EJ, Fox EE, et al. Birth order and risk of childhood cancer: a pooled analysis from five US States. Int J Cancer. 2011; 128: 2709-2716.
- Yan K, Xu X, Liu X, Wang X, Hua S, Wang C, et al. The associations between maternal factors during pregnancy and the risk of childhood acute lymphoblastic leukemia: A meta-analysis. Pediatr Blood Cancer. 2015; 62: 1162-1170.
- Schüz J, Luta G, Erdmann F, Ferro G, Bautz A, Simony SB, et al. Birth order and risk of childhood cancer in the Danish birth cohort of 1973-2010. Cancer Causes Control. 2015; 26: 1575-1582.
- Crump C, Sundquist J, Sieh W, Winkleby MA, Sundquist K. Perinatal risk factors for acute myeloid leukemia. Eur J Epidemiol. 2015; 30: 1277-1285.
- Mensah FK, Willett EV, Simpson J, Smith AG, Roman E. Birth order and sibship size: evaluation of the role of selection bias in a case-control study of non-Hodgkin's lymphoma. Am J Epidemiol. 2007; 166: 717-723.
- Erdmann F, Winther JF, Dalton SO, Lightfoot T, Zeeb H, Simony SB, et al. Survival From Childhood Hematological Malignancies in Denmark: Is Survival Related to Family Characteristics? Pediatr Blood Cancer. 2016; 63: 1096-1104.
- Tanaka A, Lindor K, Ansari A, Gershwin ME. Fetal microchimerisms in the mother: immunologic implications. Liver Transpl. 2000; 6: 138-143.
- Nelson JL. Microchimerism in human health and disease. Autoimmunity. 2003; 36: 5-9.
- Ichinohe T, Teshima T, Matsuoka K, Maruya E, Saji H. Fetal-maternal microchimerism: impact on hematopoietic stem cell transplantation. Curr Opin Immunol. 2005; 17: 546-552.
- Dawe GS, Tan XW, Xiao ZC. Cell migration from baby to mother. Cell Adh Migr. 2007; 1: 19-27.
- van Rood JJ, Scaradavou A, Stevens CE. Indirect evidence that maternal microchimerism in cord blood mediates a graft-versus-leukemia effect in cord blood transplantation. Proc Natl Acad Sci U S A. 2012; 109: 2509-2514.
- Amiel A, Leopold L, Gronich N, Yukla M, Fejgin MD, Lishner M. The influence of different chromosomal aberrations on molecular cytogenetic parameters in chronic lymphocytic leukemia. Cancer Genet Cytogenet 2006; 167: 145-149.
- Haig D, Grafen A. Genetic scrambling as a defence against meiotic drive. J Theor Biol. 1991; 153: 531-558.
- Haig D. Genomic imprinting and kinship: how good is the evidence? Annu Rev Genet. 2004; 38: 553-585.
- Ubeda F, Haig D. On the evolutionary stability of Mendelian segregation. Genetics. 2005; 170: 1345-1357.
- Ferguson-Smith AC. Genomic imprinting: the emergence of an epigenetic paradigm. Nat Rev Genet. 2011; 12: 565-575.
- Smallwood SA, Kelsey G. De novo DNA methylation: a germ cell perspective. Trends Genet. 2012; 28: 33-42.
- Hartl DL. Modifier theory and meiotic drive. Theor Popul Biol. 1975; 7: 168-174.
- Liberman U. Modifier theory of meiotic drive: is Mendelian segregation stable? Theor Popul Biol. 1976; 10: 127-132.
- Haig D. Retroviruses and the placenta. Curr Biol. 2012; 22: R609-613.
- Jønsson V, Awan H, Johannesen TB, Tjønnfjord GE. Chronic lymphocytic leukemia, advantage of monoclones?J Leuk. 2014; 2: 142-46.doi: .417102/2329-6917.1000142.
- Kennedy AE, Kamdar KY, Lupo PJ, Okcu MF, Scheurer ME, Dorak MT . Birth Characteristics and Childhood Leukemia Risk: Correlations With Genetic Markers. J Pediatr Hematol Oncol. 2015; 37: e301-307.
- O'Neill KA, Murphy MF, Bunch KJ, Puumala SE, Carozza SE, Chow EJ, et al. Infant birthweight and risk of childhood cancer: international population-based case control studies of 40 000 cases. Int J Epidemiol. 2015; 44: 153-168.
- Reigstad MM, Larsen IK, Myklebust TA, Robsahm TE, Oldereid NB, Brinton LA, et al. Risk of cancer in children born after assisted reproductive technology in Norway. Hum Reprod. 2015; 30: i17-i18.doi:10.1093/humrep/30.Supplement_1.1.
- Goldin LR, Slager SL, Caporaso NE. Familial chronic lymphocytic leukemia. Curr Opin Hematol. 2010; 17: 350-355.
- Haldane JB, Smith CA. A simple exact test for birth-order effect. Ann Eugen. 1948; 14: 117-124.
- Wilcoxin F. Probability tables for individual comparisons by ranking methods. Biometrics. 1947; 3: 119-122.