Research Article
Over-Expression of Cofilin-1 Suppressed Mobility of Lung Cancer Cells is Associated with Down-Regulation of SNAIL-1 and Induction of Let-7
Wang CY1, Tsai CH2, Chang CY2, Liao MJ2, Liu RS2-4 and Lee YJ2,5*
1Department of Medical Imaging, Cheng Hsin General Hospital, Taiwan
2Departments of Biomedical Imaging and Radiological Sciences, National Yang-Ming University, Taiwan
3Department of Nuclear Medicine, National PET/Cyclotron Center, Taiwan
4Molecular and Genetic Imaging Core, Medical School, National Yang-Ming University, Taiwan
5Biophotonics & Molecular Imaging Research Center (BMIRC), National Yang-Ming University, Taiwan
*Corresponding author: Yi-Jang Lee, Department of Biomedical Imaging and Radiological Sciences, National Yang-Ming University, No. 155, Sec. 2, Linong St. Beitou District, 112, Taipei, Taiwan
Published: 01 Jun, 2016
Cite this article as: Wang CY, Tsai CH, Chang CY, Liao
MJ, Liu RS, Lee YJ. Over-Expression of
Cofilin-1 Suppressed Mobility of Lung
Cancer Cells is Associated with DownRegulation
of SNAIL-1 and Induction of
Let-7. Clin Oncol. 2016; 1: 1015.
Abstract
Metastatic lung cancer means the spread of cancer from the primary site to nearby structures or distant organs. Epithelial-mesenchymal transition (EMT) is an important mechanism to be associated with metastasis. Suppression of EMT may prevent the cancer metastasis. We previously found that over-expression of cofilin-1, an actin binding protein belongs to the actin depolymerizing factor (ADF)/cofilin family leads to morphological change and inhibition of invasion of human non-small cells lung cancer (NSCLC). This effect is associated with up-regulation of the tumor suppressive let-7 microRNA through TWIST-1 transcription factor, an important biomarker of EMT. Here we investigated whether other EMT related molecules would be affected by overexpressed cofilin-1. Over-expression of cofilin-1 in human H1299 lung cancer cells also suppressed SNAIL-1 transcription factors, but E-cadherin and N-cadherin were not significantly affected. Importantly, over-expression of cofilin-1 induced let-7 could be suppressed by enforced expression of SNAIL-1, suggesting that EMT related transcription factors can be suppressed by over-expressed cofilin-1 to induce let-7 expression. However, over-expression of cofilin-1 may not suppress EMT. To monitor the effects of cofilin-1 and let-7 on lung cancer migration in vivo, we established a multiple reporter genes transduced lung cancer cell line that can be detected using the reporter gene imaging. The cofilin-1 induced let-7 was suppressed by transfection of locked nucleic acid (LNA) to inhibit let-7. Compared to normal lung cancer cells, over-expression of cofilin-1 suppressed the lung cancer migration, but simultaneously transfection of LNA recovered their migration ability to lungs in small animals. Taken together, over-expression of cofilin-1 can suppress the invasion and migration of lung cancer cells through up-regulation of let-7 in vitro and in vivo. Additionally, cofilin-1 may regulate EMT related transcription factors but not the whole EMT mechanism.
Keywords: Cofilin-1; SNAIL-1; let-7; EMT; Reporter gene imaging; Locked nucleic acid
Introduction
According to American Cancer Society's estimation, lung cancer is the second most common
cancer type independent of sex. However, lung cancer is the leading cause of death in cancer patients
worldwide [1]. More than 85% of lung cancer belongs to non-small cells lung cancer (NSCLC) that
includes several subtypes, such as adenocarcinoma, squamous cell carcinoma, large carcinoma and
less commonly found adenosquamous carcinoma. Metastasis is the primary cause of lung cancer
death, including bone and brain metastasis. Therefore, suppression of metastasis is important for
tumor control.
Actin cytoskeleton is formed by Rho small GTPase signaling pathway that can form different
types of actin architectures for cell morphology, attachment and migration. This signaling mediates
the activity of cofilin-1, a non-muscle is form of actin depolymerizing factor (ADF)/cofilin family
member to accelerate the actin dynamics [2,3]. Over-expression of cofilin-1 may disrupt the balance
of actin dynamics and lead to obstacle of cell motility. However, the underlying mechanisms remain
to be addressed.
Metastasis is strongly associated with the epithelial –mesenchymal
transition (EMT). EMT is a process of cell morphological change that
allows cancer penetrating through the vessel and traveling to distant
organs for regrowth [4,5]. Several markers of EMT with the property
of transcription factors have been widely reported, including Twist
Basic Helix-Loop-Helix Transcription Factor 1 (TWIST-1), Zinc
finger protein SNAI1 (SNAIL-1) and SNAI2 (Slug), Zinc finger
E-box-binding homeobox 1/2 (ZEB1 and ZEB2) [6,7]. E-cadherin is
responsible for cell-cell adhesion and is usually down-regulated by
these transcription factors followed by the emergence of mesenchymal
shapes [8]. N-cadherin is up-regulated to balance the down-regulated
E-cadherin for altered cell adhesion [9]. These transcription
factors also promote anti-apoptosis, angiogenesis, chromosomal
instability, and are generally regarded oncogenes. Interestingly,
recent reports showed that TWIST-1 can suppress the expression
of tumor suppressive lethal-7 (let-7) microRNA, which can further
regulate Ras and high mobility group A2 (HMGA2) oncogenes
[10-12]. Additionally, low let-7 level significantly correlates to the
postoperative death [13]. However, it is unclear whether different
EMT-related markers will influence the expression of let-7 or not.
Because over-expression of cofilin-1 would disrupt the actin
dynamics, it is speculated that cell migration should be affected.
Previously, we have found that over-expressed cofilin-1 could
suppress the expression of TWIST-1 in NSCLC cells [14]. The let-
7 family members were subsequently up-regulated, and let-7b and
let-7e exhibited most significant up-regulation. Because TWIST-1 is
involved in EMT, it is interesting to investigate whether other EMT
related molecules can regulate the expression of let-7 microRNA.
In this study we further examined the expression of SNAIL-1 and
E-cadherin after over-expression of cofilin-1 in lung cancer cells.
Additionally, we used reporter gene imaging to monitor whether
knockdown of let-7 in cofilin-1 over-expressing lung cancer cells
would recover the migration ability in small animals. This study
would establish a signaling pathway between cofilin-1 and let-7 that
regulate lung cancer migration in vitro and in vivo.
Figure 1
Figure 1
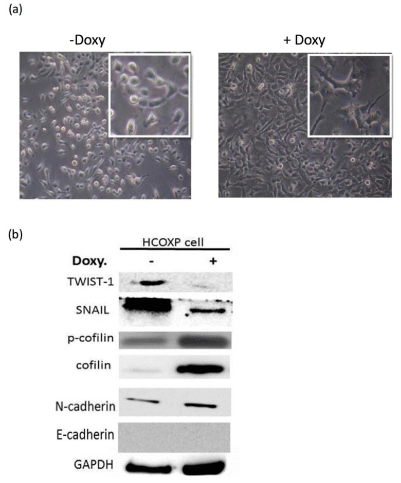
Effects of over-expressed cofilin-1 on EMT related molecules. (a) Morphological change induced by over-expression of cofilin-1. Insets are enlarged
photos of normal cells and cofilin-1 over-expressing cells. (b) Western blot analysis for detection of EMT related molecules in response to over-expression of cofilin-1.
Materials and Methods
Cell culture
Human lung cancer H1299 cells with tet-on inducible cofilin-1
over-expression cell line (HCOXP) and reporter genes harboring
HCOXP cells (HCOXP-3R) were maintained in Dulbecco's Modified
Eagle's medium (DMEM) supplemented with 10% fetal bovine
serum, 1% penicillin-streptomycin solution, 2mM L-glutamine
(Sigma-Aldrich Co, St. Louis, MO, USA), and 0.1mg/ml hygromycine
B (Invitrogen, Carlsbad, CA, USA). HCOXP cells were maintained
in a humidified incubator with 5% CO2 and 37ºC by passaged every 48 hours.
Western blot analysis and antibodies
The procedure of protein extraction, gel running and electrotransferring
has been described previously [15]. The primary
antibodies used in this study include: anti-cofilin-1, anti- ser3-
phospho-cofilin-1, anti-Twist-1, anti-TWIST-1 (Genetex Inc.,
Irvine, CA, USA), anti-SNAIL-1, anti-cofilin-1 (Genetex Inc., Irvine,
CA, USA), anti-phospho-cofilin-1 (Santa Cruz Biotechnology Inc.,
Dallas, TX, USA), anti-E-cadherin, anti-N-cadherin, anti-HSV1-tk
(Santa Cruz Biotechnology Inc., Dallas, TX, USA) and anti- GAPDH
(Sigma-Aldrich Co, St. Louis, MO, USA) antibodies.
Quantification of let-7
To measure let-7 microRNA levels before and after cofilin-1
over-expression, quantitative PCR (qPCR) of targeted miRNA was
used. In brief, complementary DNA (cDNA) was generated from
5 µg total RNA using SuperScript II reverse transcriptase (LifeTechnologies
Co, Carlsbad, CA, USA). The cDNA products were
then mixed in the Fast SYBR Green Master Mix (Life-Technologies
Co, Carlsbad, CA, USA) and subjected to the Step One Plus RealTime
PCR System (Life-Technologies Co, Carlsbad, CA, USA)
according to the manufacturer's instructions. The stem loop
primers used for let-7b and let-7e were 5'-GTCGTATCCAGTG
CAGGGTCCGAGGTATTCGCACTGGATACGACAACcac-3' and
5'-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGC ACTGGAT
ACGACAACTAT-3', respectively. The forward primers of qPCR used
for let-7b and let-7e were 5'- GCCGCTTGAGGTAGTAGGTTGT-3'
and 5'-GCCGCTTGAGGtAGGAGGTTGT-3', respectively. The
universal reverse primer was used for both let-7: 5'-CCAGTGC
AGGGTCCGAGGT-3'. For internal control, the vertebrate U6 small
nuclear RNA was amplified using the primer set: 5'-CGCTTCGGC
AGCACATATAC-3' and 5'-TTCACGAATTTGCGTGTCAT-3'.
Establishment of HCOXP-3R cells and reporter gene validations –
LT-3R plasmid, a multicistronic lentiviral construct was used to
establish HCOXP-3R cells for expressing firefly luciferase (fLuc),
green fluorescent protein (GFP) and herpes simplex virus type
1-thymidine kinase (HSV1-tk) reporter genes [16]. This plasmid was
co-transfected with pCMV-∆R8.91 plasmid and pMD.G plasmid into
the 293T packaging cell line to produce virion soup using the calcium
phosphate precipitation method. Ultracentrifugation was used to
concentrate the virion soup, which was added to HCOXP cell culture
for infection. After infection, cells were sorted by the fluorescenceactivated
cell sorting (FACS, FACSAria, BD Biosciences, San
Jose, CA, USA) based on the GFP emitted fluorescent signals. The
obtained stable clone was named HCOXP-3R cells, in which the GFP
expression was visualized using the fluorescent microscope. The fLuc
activity and expression of HSV1-tk protein were determined by the
luciferase assay and Western blot analysis as described before [16].
Knockdown of let-7 – Chemically modified locked nucleic acid
(LNATM, Exiqon, Los Angeles, CA, USA) was used to silence the
expression of let-7 microRNA. In brief, 30nM LNA was transfected
into HCOXP-3R cells using the JetPEI transfection reagent (Polyplustransfection,
SA, Illkirch, France). The sequences of LNA for targeting
on let-7b and let-7e were 5'-ACCACACAACCTACTACCTC-3' and
5'-ACTATACAACCTCCTACCTC-3', respectively.
Luciferase assay – The expression of luciferase activity in cells with
over-expressed cofilin-1 and co-transfected LNA were determined
using the in vitro luciferase assay. Cells were cultured in 12-well plates
and lyzed by passive lysis buffer and then added with 5-fold diluted
reporter assay buffer (50mM glycylglycin, 1M magnesium sulfate,
10mg/ml bovine serum albumin, and 0.5M EDTA) mixed with
100mM adenosine 5'-triphosphate disodium salt (Sigma-Aldrich
Co., St. Louis, MO, USA), 1M dithiothreitol and 50mM D-luciferin
luciferin (Promega Co., Madison, WI, USA) transferring to a 96-wel
black plate. The luminescent signals were detected using a multimode
microplate reader (TECAN, Switzerland).
In vitro invasion assay – Cells were trypsinized and five thousand
cells were seeded in transwells coated with Matrigel (BD biosciences,
San Jose, CA, USA) in serum-free DMEM. Each transwell was placed
in a 24-well dish containing DMEM with 10% FBS. After 48 hours of
incubation, the transwells were cleaned with a cotton stub, fixed using
4% paraformaldehyde followed by crystal violate (1.25% in ethanol)
staining for 30 minutes. The transwells were then rinsed, visualized
and counted under a bright-field microscope.
Experimental metastasis animal model and bioluminescent imaging
HCOXP-3R cells (1x106) were injected into the nude mice via
tail veins. After injection, the animals were subjected to IVIS-50
imaging system (Caliper Co, Hopkinton, MA, USA) to detect the
bioluminescent signals in vivo. Before imaging, the animals were i.p.
injected with 150mg/kg D-luciferin (VivoGlo Luciferin, Promega
Corp., Madison, WI, USA) and anesthetized with 1% isofluorane for
15 minutes. The images were acquired using the bundled software.
The animal studies have been approved by Institutional Animal Care
and Use Committee (IUCAC No. 1021208) of National Yang-Ming
University.
Statistic analysis
Each datum represented means ± S.D., and the results were
analyzed by student's t-test between two samples. For multiple
samples, one-way analysis of variance (ANOVA) was used for statistic
analysis. In both conditions, p < 0.05 was regarded significance.
The analysis and plots were executed using Sigmaplot 10.0 software
(Systat Software, Inc, a Jose, CA, USA).
Results
Effects of cofilin-1 over-expression on EMT related molecules
HCOXP cells are derived from H1299 lung cancer cells harboring
a tet-on gene expression system for over-expression of cofilin-1.
Because induction of cofilin-1 expression in these cells will lead
to apparent morphological change likes EMT, we investigated
whether over-expression of cofilin-1 would influence the EMT
related molecules (Figure 1A). TWIST-1, SNAIL-1, N-cadherin and
E-cadherin were examined after over-expression of cofilin-1 using the
Western blot analysis. Although TWIST-1 and SNAIL-1 were downregulated
by over-expressed cofilin-1, E-cadherin and N-cadherin
levels were not changed significantly (Figure 1B). Over-expressed
cofilin-1 can be phosphorylated on the serine-3 as described before.
Therefore, these data suggest that cofilin-1 would influence the
expression of TWIST-1 and SNAIL-1 transcription factors but not
the whole EMT related biomarkers.
Over-expression of cofilin-1 induced let-7 up-regulation
was suppressed by SNAIL-1
Previously, we have found that over-expression of cofilin-1 can
induce let-7 microRNA through suppression of TWIST-1 expression
[14]. Over-expression of TWIST-1 can counteract the induced let-
7 microRNA by over-expressed cofilin-1. Here we investigated
whether SNAIL-1 transcription factor can also regulate let-7 or not.
In HCOXP cells, induction of cofilin-1 expression could induce let-
7b and let-7e that have been reported to be most responsive to overexpressed
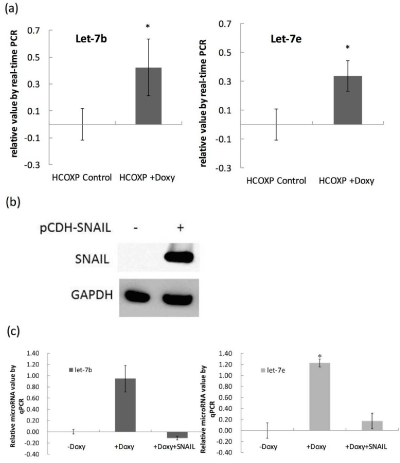
cofilin-1 (Figure 2A). We next showed that in doxycycline
treated HCOXP cells, transfection of pCDH-SNAIL-1 construct could
restore the SNAIL-1 level in these cells (Figure 2B). This treatment
led to suppression of let-7b and let-7e induced by over-expressed
cofilin-1 (Figure 2C). Therefore, down-regulation of SNAIL-1 by
over-expressed cofilin-1 is also involved in regulate the expression of
let-7 microRNA.
Characterization of HCOXP-3R cells responding to knockdown
of let-7 after over-expression of cofilin-1 – We have previously
established a lentiviral-based multicistronic reporter construct
including green fluorescent protein gene (GFP), firefly luciferase
(fLuc), and herpes virus type 1 – thymidine kinase (HSV1-tk)
gene, which are used for examination of transfection efficiency,
bioluminescent imaging, and radionuclide based imaging in vivo,
respectively. Here we transduced this construct into HCOXP cells
that have not been examined before. The obtained stable cells were
named HCOXP-3R cells. The expression of GFP was examined using
the fluorescent microscope (Figure 3A). The activity of fLuc was
determined using the luciferase assay with luciferin substrate (Figure
3B). The expression of HSV1-tk was examined using Western blot
analysis (Figure 3C). This novel stable lung cancer cell line would
be used for in vivo tracking under different conditions. We next
compared the cell viability and invasive ability in HCOXP-3R cells
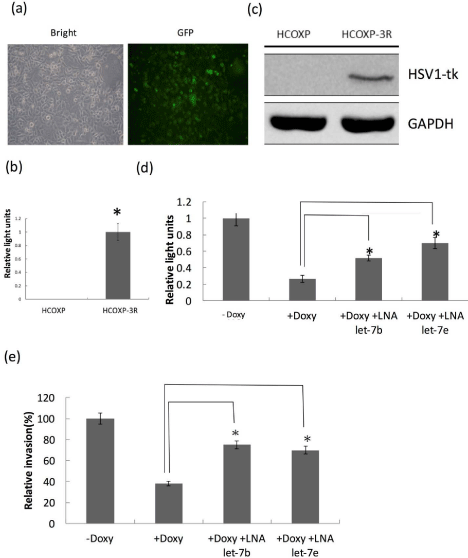
before and after over-expression of cofilin-1. The luciferase activity
of HCOXP-3R cells were inhibited by induced cofilin-1 expression,
but co-treatment of LNA also recovered the luciferase activity
(Figure 3D). HCOXP-3R cells invading through the matrigel coated
transwells were further compared by above conditions. The results
showed that over-expression of cofilin-1 could suppress the invasion,
but the effects were compromised after treatment of let-7 targeting
LNA (Figure 3E). Thus, the in vivo effect of over-expressed cofilin-1
on lung cancer was subsequently examined using this novel stable
clone.
Bioluminescent imaging of lung accumulation by HCOXP-3R
cells responding to cofilin-1 over-expression and LNA treatment –
We next investigated whether migration of HCOXP-3R cells to lungs
will be affected by over-expressed cofilin-1 and co-treated LNA.
HCOXP-3R cells were either treated with doxycycline to induce
cofilin-1 expression or transfected with LNAs followed by doxycycline
treatment. These cells (1x106 each) were then separately i.v. injected
into nude mice. After injection, the mice were subjected to IVIS-
50 system and imaged for the bioluminescent signals immediately.
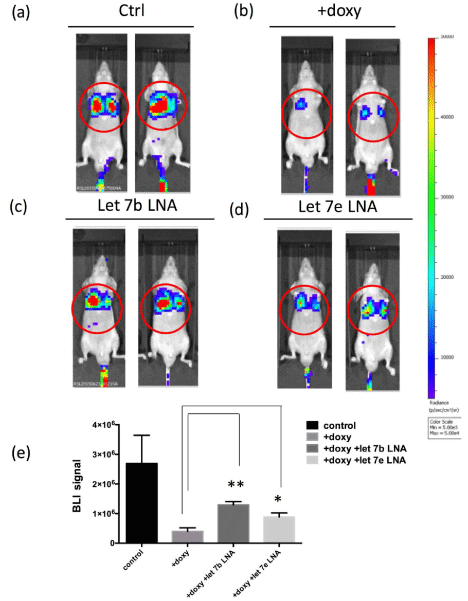
Compared to untreated cells, over-expression of cofilin-1 apparently
inhibited the migration of HCOXP-3R cells to lungs (Figure 4A and B).
However, suppression of let-7b or let-7e by LNAs could recover
the lung migration of cofilin-1 over-expressing cells (Figure 4C and
D). The photons flux in chest of each group was also semi-quantified
according to the analytic software of IVIS system (Figure 4E).
Therefore, the effects of cofilin-1 signaling pathways on lung cancer
growth and metastasis in vivo would be easily examined using the
HCOXP-3R cells in the future.
Figure 2
Figure 2
Effects of SNAIL-1 on suppression of let-7 induced by over-expression of cofilin-1. (a) Quantification of let-7b and let-7e using qPCR before and after over-expression of cofilin-1. (b) Transfection of pCDH-SNAIL-1 to HCOXP cells followed by Western blot analysis. (c) Over-expression of SNAIL-1 suppressed the
expression of let-7b and let-7e induced by cofilin-1.
Discussion
Previously, over-expression of cofilin-1 was found to induce let-7
microRNA in human H1299 lung cancer cells. This signaling pathway
was mediated by TWIST-1 rather than other let-7 regulators, that is,
LIN28B and c-Myc [14]. Because TWIST-1 is known to be one of
the important markers of EMT, it is of interest to investigate whether
over-expression of cofilin-1 induced let-7 is caused by suppression of
EMT. Although SNAIL-1 was also down-regulated by over-expressed
cofilin-1, another EMT markers E-cadherin and N-cadherin were
not significantly affected under this condition. The basal level
of E-cadherin in H1299 cells were barely detected as reported
previously [17], while suppression of TWIST-1 and SNAIL-1 by over
expressed cofilin-1 did not up-regulate E-cadherin. N-cadherin was
also not reduced. Therefore, the current data suggest that cofilin-1
may influence certain EMT related molecules rather than the EMT
phenomenon.
The expression of let-7 has been reported to be regulated by
LIN28A/B, c-Myc and TWIST-1 during cancer development
[12,18,19]. As a tumor suppressor, up-regulation of let-7 leads to
suppression of tumor metastasis and cancer growth. Interestingly,
a recent report showed that SNAIL-1 can temporarily bind to let-7
promoters and reduce its expression for efficient reprogramming of
fibroblasts [20]. Here we found that let-7b and let-7e induced by overexpressed
cofilin-1 could be suppressed by transfection of SNAIL-1.
This result suggests that the SNAIL-1 regulated let-7 expression also
plays a role in lung cancer cells. Like TWIST-1, SNAIL-1 also mediates
the cofilin-1 regulated let-7 expression. How over-expression of
cofilin-1 leads to down-regulation of TWIST-1 and SNAIL-1 but
not E-cadherin is unclear and may be important to be studied in the
future.
The most critical experiments in this report were the first time
to demonstrate the cofilin-1/let-7 signaling pathway would affect the
migration of lung cancer cells to lungs in vivo. To this end, we used
the reporter gene imaging to track the positions of cancer cells in
the nude mice. The multicistronic reporter gene construct, so called
LT-3R has been used in tracking the growth of glioblastomas in vivo
[16]. Here we transduced this construct to HCOXP cells, which were
subsequently i.v. injected into small animals. This experimental
metastasis model has been used for investigating the cancer cells
migration to lungs in different cancer types [21,22]. The LT-3R
plasmid transduced HCOXP cells exhibited similar phenotypes
with parental HCOXP cells, including cell growth and growth
suppression by over-expressed cofilin-1. Because reporter genes only
express in viable cells [23], the luciferase assay should be sufficient
to demonstrate the effects of over-expressed cofilin-1 on suppression
of cell viability. Transfection of let-7 targeted LNA in cofilin-1 overexpressing
HCOXP-3R cells led to recovery of luciferase expression,
suggesting that let-7 is important for mediating the effects of cofilin-1
on cell growth and viability. Furthermore, the cellular invasive ability
suppressed by over-expressed cofilin-1 was also recovered by LNA.
These in vitro studies support that HCOXP-3R cells can used as a
surrogate to monitor the behaviors of HCOXP cells in vivo.
To investigate the cancer metastasis in small animals, both
"experimental metastasis" (intravascular injection of cells) and
"spontaneous metastasis" (orthotopic injection of cells) are usually
applied [24]. Because lung cancer model was used here, we first
examined whether over-expression of cofilin-1 could affect the
migration of these cells to lungs via i.v. injection, and whether LNA
would reverse the cofilin-1 effects as found in in vitro studies. The data
showed that the in vivo study was consistent with in vitro effects of
cofilin-1 over-expression. However, we could not detect the migration
of HCOXP-3R cells out of lungs because the bioluminescent signals
were disappeared on next day (data not shown). This should not
due to the loss of reporter genes because the expression of reporter
genes in HCOXP-3R cells remains detectable after one month of
continuous culturing. The current data suggest that over-expression
of cofilin-1 may inhibit the migration of HCOXP-3R cells to lungs,
but suppression of let-7 in these cells will compromise this effect.
Whether different levels of bioluminescent signals detected in lungs
represents reduced extravasation of HCOXP -3R cells after overexpression
of cofilin-1 is of interest to further investigate.
In summary, this is the first study showing that over-expressed
cofilin-1 can inhibit the migration of lung cancer in vivo, to the best
of our knowledge. We also demonstrate that cofilin-1 induced let-7 is
required for inhibition of migration because knockdown of let-7 by
LNA can reverse this effect. Although down-regulation of TWIST-1
has been previously reported to be required for cofilin-1 mediated
up-regulation of let-7, SNAIL-1 transcription factor is also involved
in this signaling pathway. However, it appears that EMT is not the
primary role to be ablated by over-expressed cofilin-1. Taken together,
the in vivo evidence of cofilin-1/let-7 pathway on controlling the lung
cancer migration would be important for clinical consideration of
molecular targeting therapy.
Figure 3
Figure 3
Effects of knockdown of let-7 in HCOXP cells with co-expressed reporter genes. (a) Fluorescent microscope for visualizing GFP. (b) Luciferase assay for detecting the luciferase activity. (c) Detection of HSV1-tk protein by Western blot analysis. (d) Comparison of cell viability by detecting the luciferase activity in
cofilin-1 over-expressing cells before and after transfection of LNA. (e) Comparison of in vitro invasion ability according to above conditions. *: p<0.05.
Figure 4
Figure 4
In vivo imaging of HCOXP-3R cells distribution through intravenous injection into nude mice. (a) Untreated HCOXP-3R cells. (b) Doxycycline treated HCOXP-3R cells. (c) Let-7b and (d) let-7e targeted LNA transfected HCOXP-3R cells that have been treated with doxycycline. (e) The regions of interest (ROI) of
each group were marked by red circles and quantified for the BLI signals. *: p < 0.05. **: p < 0.01 (N=4).
Acknowledgement
This study was supported by a united grant of Cheng-Hsin General Hospital and National Yang-Ming University (103F003C05), and the Ministry of Science and Technology of Taiwan (102-2628-B-010-012-MY3).
References
- Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61: 69-90.
- Carlier MF, Laurent V, Santolini J, Melki R, Didry D, Xia GX, et al. Actin depolymerizing factor (ADF/cofilin) enhances the rate of filament turnover: implication in actin-based motility. J Cell Biol. 1997; 136: 1307- 1322.
- Mizuno K. Signaling mechanisms and functional roles of cofilin phosphorylation and dephosphorylation. Cell Signal. 2013; 25: 457-469.
- Lee JM, Dedhar S, Kalluri R, Thompson EW. The epithelial-mesenchymal transition: new insights in signaling, development, and disease. J Cell Biol. 2006; 172: 973-981.
- Popper HH. Progression and metastasis of lung cancer. Cancer Metastasis Rev. 2016; 35: 75-91.
- Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelialmesenchymal transition. Nat Rev Mol Cell Biol. 2014; 15: 178-196.
- Zeisberg M, Neilson EG. Biomarkers for epithelial-mesenchymal transitions. J Clin Invest. 2009; 119: 1429-1437.
- Yilmaz M, Christofori G. EMT, the cytoskeleton, and cancer cell invasion. Cancer Metastasis Rev. 2009; 28: 15-33.
- Wheelock MJ, Shintani Y, Maeda M, Fukumoto Y, Johnson KR. Cadherin switching. J Cell Sci. 2008; 121: 727-735.
- Lee YS, Dutta A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007; 21: 1025-1030.
- Wang X, Cao L, Wang Y, Wang X, Liu N, You Y. Regulation of let-7 and its target oncogenes (Review). Oncol Lett. 2012; 3: 955-960.
- Yang WH, Lan HY, Huang CH, Tai SK, Tzeng CH, Kao SY, et al. RAC1 activation mediates Twist1-induced cancer cell migration. Nat Cell Biol. 2012; 14: 366-374.
- Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res. 2004; 64: 3753-3756.
- Tsai CH, Lin LT, Wang CY, Chiu YW, Chou YT, Chiu SJ, et al. Overexpression of cofilin-1 suppressed growth and invasion of cancer cells is associated with up-regulation of let-7 microRNA. Biochim Biophys Acta. 2015; 1852: 851-861.
- Lee YJ, Sheu TJ, Keng PC. Enhancement of radiosensitivity in H1299 cancer cells by actin-associated protein cofilin. Biochem Biophys Res Commun. 2005; 335: 286-291.
- Lin LT, Chiou SH, Lee TW, Liu RS, Hwang JJ, Chang CH, et al. A comparative study of primary and recurrent human glioblastoma multiforme using the small animal imaging and molecular expressive profiles. Mol Imaging Biol. 2013; 15: 262-272.
- Asnaghi L1, Vass WC, Quadri R, Day PM, Qian X, Braverman R, et al. E-cadherin negatively regulates neoplastic growth in non-small cell lung cancer: role of Rho GTPases. Oncogene. 2010; 29: 2760-2771.
- Chang TC, Zeitels LR, Hwang HW, Chivukula RR, Wentzel EA, Dews M, et al. JT: Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation. Proc Natl Acad Sci U S A. 2009; 106: 3384-3389.
- Wang T, Wang G, Hao D, Liu X, Wang D, Ning N, et al. Aberrant regulation of the LIN28A/LIN28B and let-7 loop in human malignant tumors and its effects on the hallmarks of cancer. Mol Cancer. 2015; 14: 125.
- Unternaehrer JJ, Zhao R, Kim K, Cesana M, Powers JT, Ratanasirintrawoot S, et al. The epithelial-mesenchymal transition factor SNAIL paradoxically enhances reprogramming. Stem Cell Reports. 2014; 3: 691-698.
- Kjønniksen I, Storeng R, Pihl A, McLemore TL, Fodstad O. A human tumor lung metastasis model in athymic nude rats. Cancer Res. 1989; 49: 5148-5152.
- Rashid OM, Nagahashi M, Ramachandran S, Dumur CI, Schaum JC, Yamada A, et al. Is tail vein injection a relevant breast cancer lung metastasis model? J Thorac Dis. 2013; 5: 385-392.
- Shin JH, Chung JK, Kang JH, Lee YJ, Kim KI, So Y, et al. Noninvasive imaging for monitoring of viable cancer cells using a dual-imaging reporter gene. J Nucl Med .2004; 45: 2109-2115.
- Price JE. Spontaneous experimental metastasis models: nude mice. Methods Mol Biol. 2014; 1070: 223-233.